Publications¶
Preprints¶
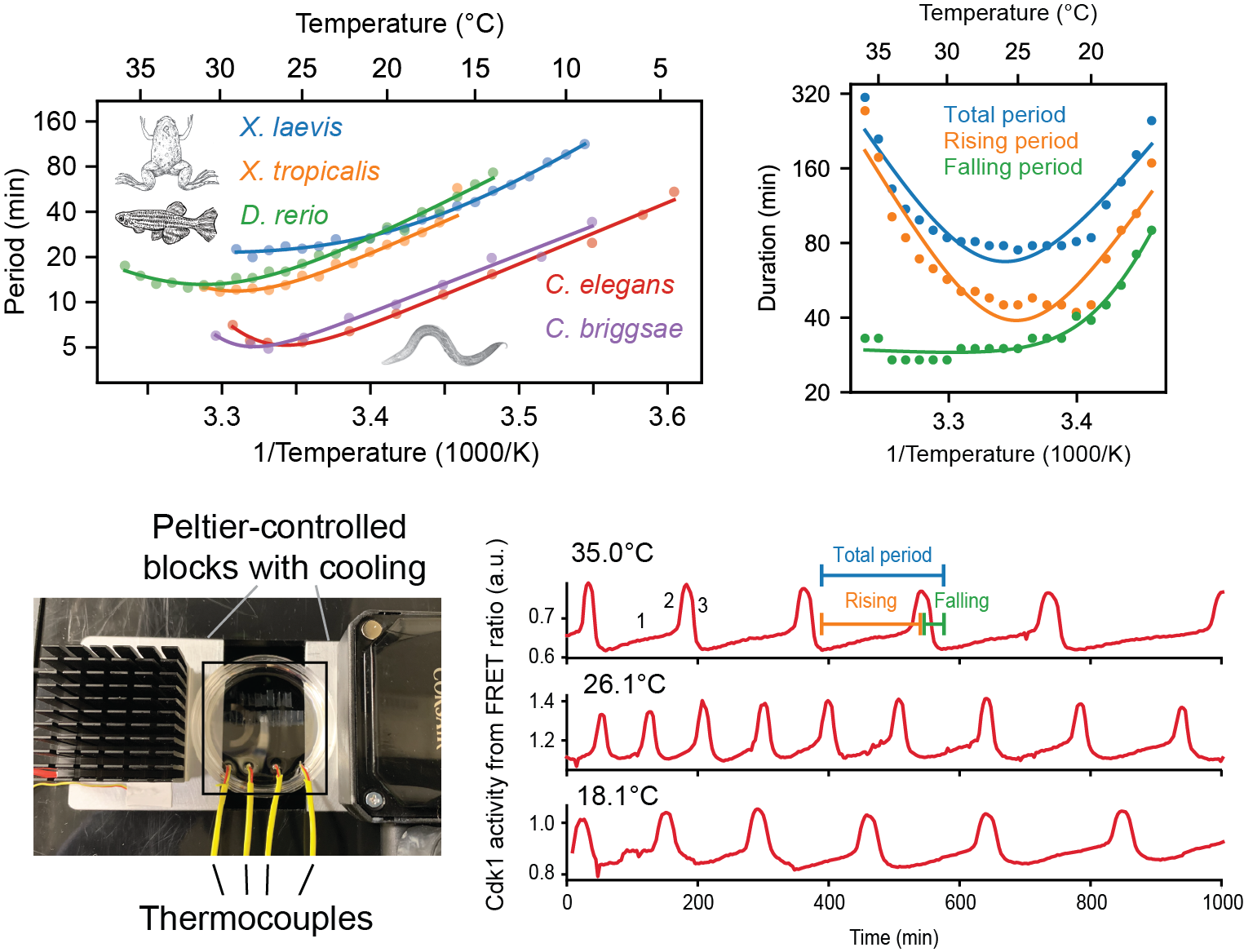
Mechanistic origins of temperature scaling in the early embryonic cell cycle
J. Rombouts*, F.Tavella*, A. Vandervelde, C. Phong, J. Ferrell, Q. Yang, L. Gelens
BioRxiv (2025)
DOI: 10.1101/2024.12.24.630245 | Link
J. Rombouts*, F.Tavella*, A. Vandervelde, C. Phong, J. Ferrell, Q. Yang, L. Gelens
BioRxiv (2025)
DOI: 10.1101/2024.12.24.630245 | Link
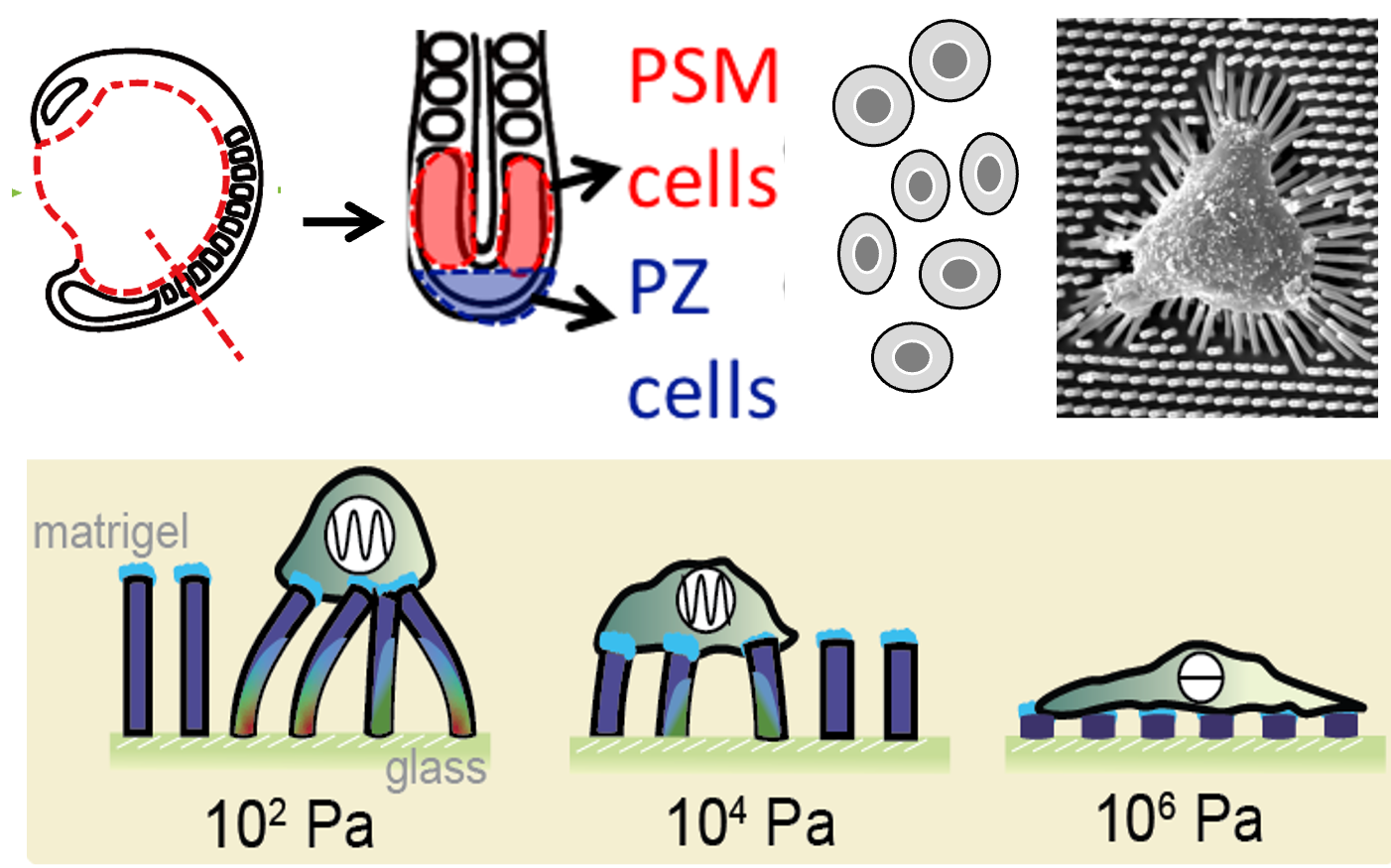
Substrate rigidity modulates segmentation clock dynamics in isolated presomitic mesoderm cells
C.-Y. Sung*, U. Kadiyala*, O. Blanchard*, L. Yourston, D. Walker, L. Li, J. Fu, Q. Yang
BioRxiv (2024)
DOI: 10.1101/2024.07.02.601712 | Link
C.-Y. Sung*, U. Kadiyala*, O. Blanchard*, L. Yourston, D. Walker, L. Li, J. Fu, Q. Yang
BioRxiv (2024)
DOI: 10.1101/2024.07.02.601712 | Link
Publications¶
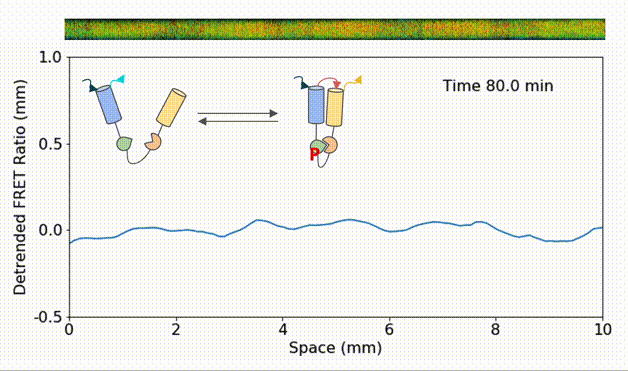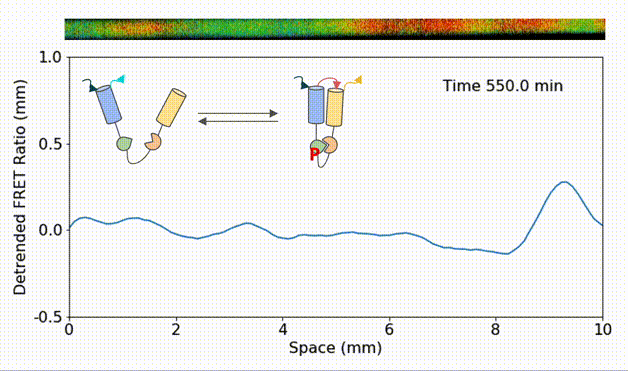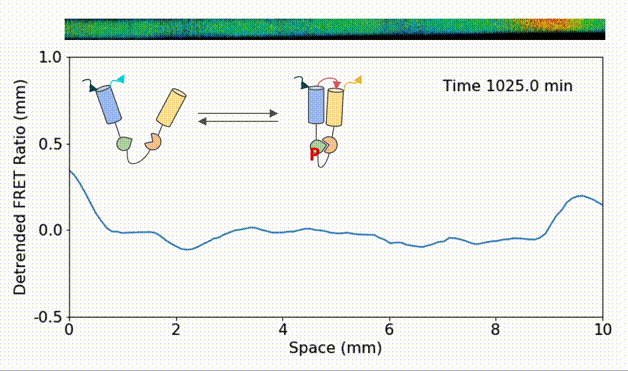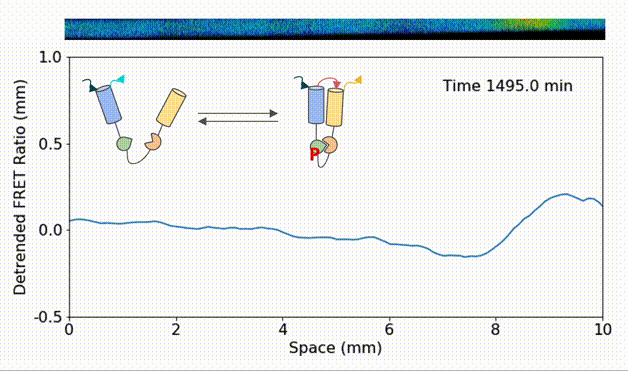
Spatial heterogeneity accelerates phase-to-trigger wave transitions in frog egg extracts
O. Puls*, D. R. Reynés*, F. Tavella, M. Jin, Y. Kim, L. Gelens, Q. Yang
Nature Communications (2024)
DOI: 10.1038/s41467-024-54752-7 | Link
O. Puls*, D. R. Reynés*, F. Tavella, M. Jin, Y. Kim, L. Gelens, Q. Yang
Nature Communications (2024)
DOI: 10.1038/s41467-024-54752-7 | Link
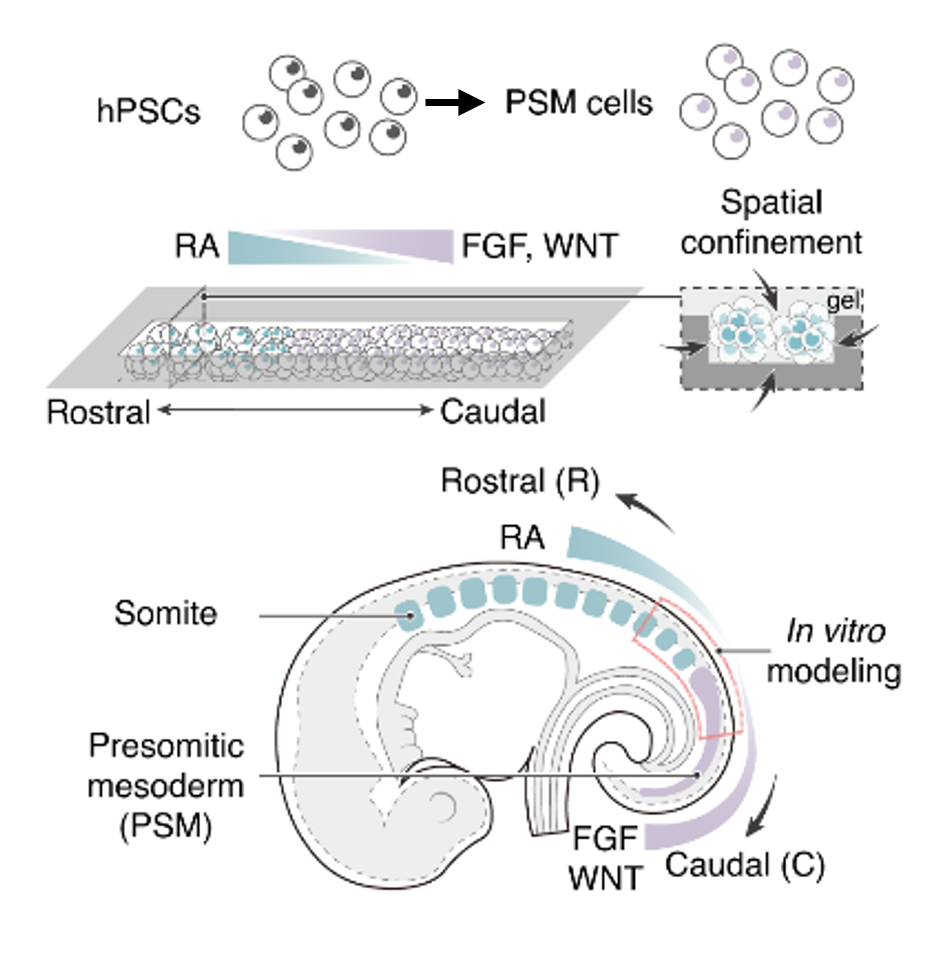
A human pluripotent stem cell-based somitogenesis model using microfluidics
Y. Liu, Y. S. Kim, X. Xue, N. Kobayashi, S. Sun, Q. Yang, O. Pourquié, J. Fu
Cell Stem Cell (2024)
DOI: 10.1016/j.stem.2024.06.004 | Link
Y. Liu, Y. S. Kim, X. Xue, N. Kobayashi, S. Sun, Q. Yang, O. Pourquié, J. Fu
Cell Stem Cell (2024)
DOI: 10.1016/j.stem.2024.06.004 | Link
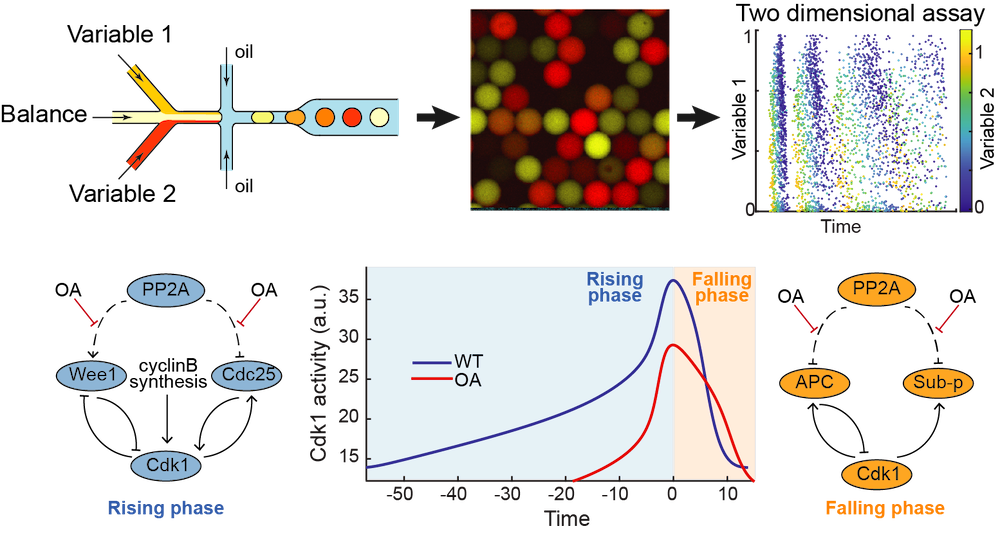
Comprehensive parameter space mapping of cell cycle dynamics under network perturbations
Z. Li*, S. Wang*, M. Sun*, M. Jin, D. Khain, Q. Yang
ACS Synthetic Biology (2024)
DOI: 10.1021/acssynbio.3c00631 | Link
Z. Li*, S. Wang*, M. Sun*, M. Jin, D. Khain, Q. Yang
ACS Synthetic Biology (2024)
DOI: 10.1021/acssynbio.3c00631 | Link
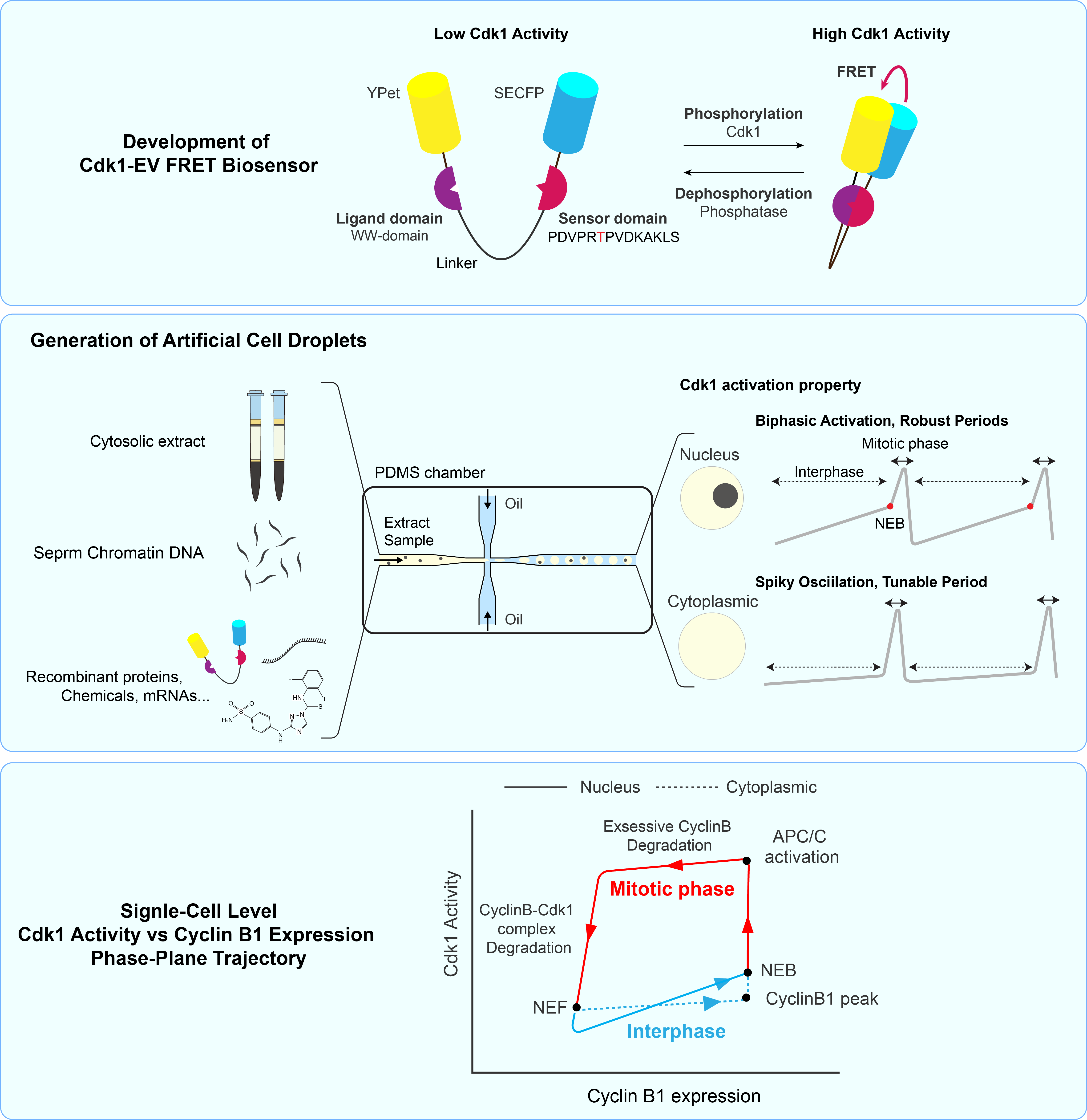
Nuclear-cytoplasmic compartmentalization of cyclin B1-Cdk1 promotes robust timing of mitotic events
G. Maryu and Q. Yang
Cell Reports (2022)
DOI: 10.1016/j.celrep.2022.111870 | Link
G. Maryu and Q. Yang
Cell Reports (2022)
DOI: 10.1016/j.celrep.2022.111870 | Link
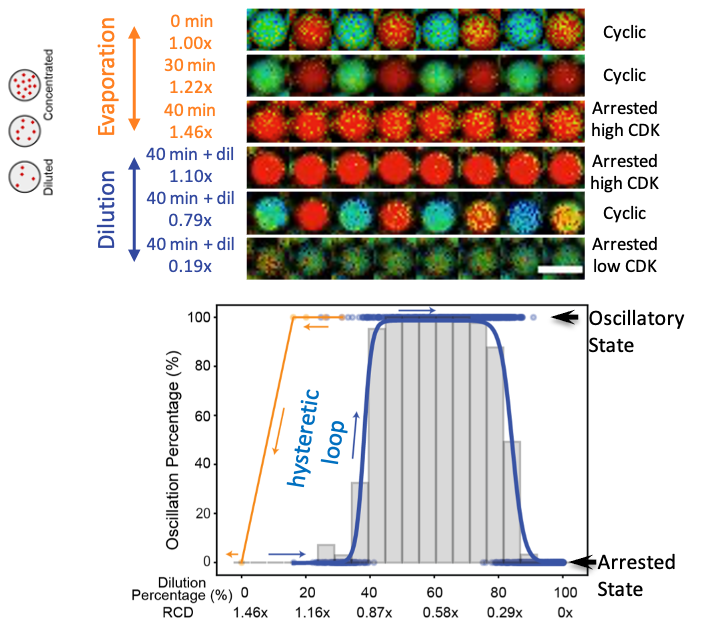
In vitro cell cycle oscillations exhibit a robust and hysteretic response to changes in cytoplasmic density
M. Jin*, F. Tavella*, S. Wang, Q. Yang
Proceedings of the National Academy of Sciences (2022)
DOI: 10.1073/pnas.2109547119 | Link
M. Jin*, F. Tavella*, S. Wang, Q. Yang
Proceedings of the National Academy of Sciences (2022)
DOI: 10.1073/pnas.2109547119 | Link
Monitoring spontaneous quiescence and asynchronous proliferation-quiescence decisions in prostate cancer cells
A. Pulianmackal, D. Sun, K. Yumoto, Z. Li, Y. Chen, M. Patel, Y. Wang, E. Yoon, A. Pearson, Q. Yang, R. Taichman, F. Cackowski, L. Buttitta
Frontiers in Cell and Developmental Biology (2021)
DOI: 10.3389/fcell.2021.728663 | Link
A. Pulianmackal, D. Sun, K. Yumoto, Z. Li, Y. Chen, M. Patel, Y. Wang, E. Yoon, A. Pearson, Q. Yang, R. Taichman, F. Cackowski, L. Buttitta
Frontiers in Cell and Developmental Biology (2021)
DOI: 10.3389/fcell.2021.728663 | Link
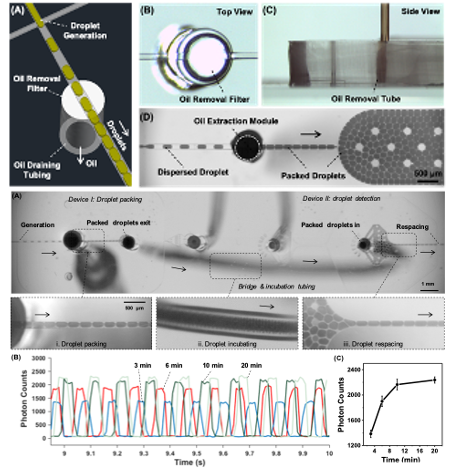
Plug-in tubes removing oil and packing droplets for time-controlled droplet-based assays
M. Sun, G. Maryu, S. Wang, Q. Yang, R. C. Bailey
Biomicrofluidics (2021)
DOI: 10.1063/5.0047924 | Link
M. Sun, G. Maryu, S. Wang, Q. Yang, R. C. Bailey
Biomicrofluidics (2021)
DOI: 10.1063/5.0047924 | Link
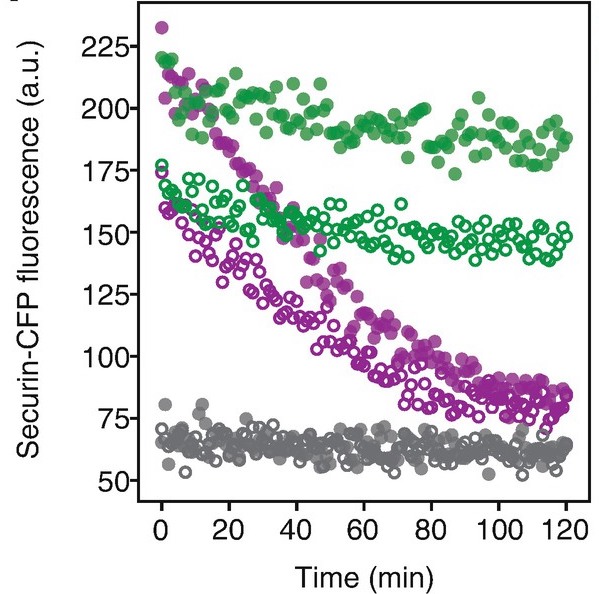
Real-time monitoring of APC/C-Mediated substrate degradation using Xenopus laevis egg extracts
J. Kamenz, R. Qiao, Q. Yang, J. E. Ferrell Jr.
Methods in Molecular Biology (2021)
DOI: 10.1007/978-1-0716-1538-6_3 | Link
J. Kamenz, R. Qiao, Q. Yang, J. E. Ferrell Jr.
Methods in Molecular Biology (2021)
DOI: 10.1007/978-1-0716-1538-6_3 | Link
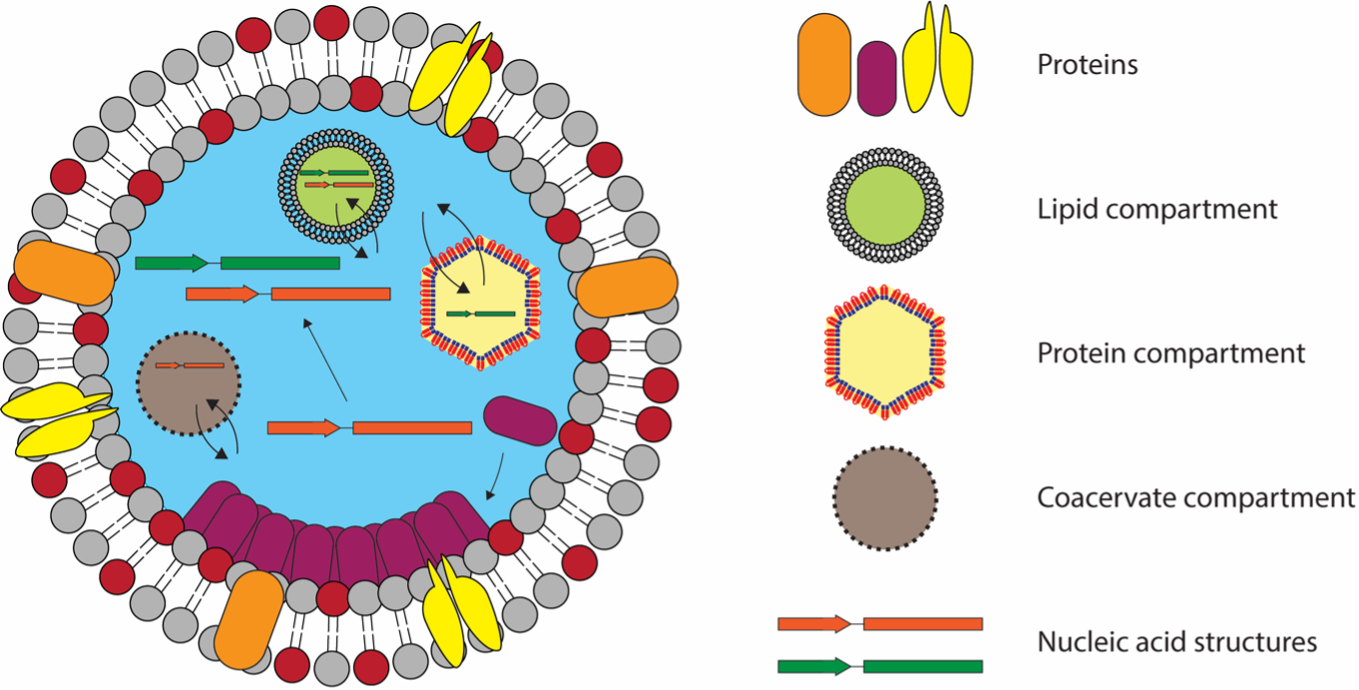
Engineering spatiotemporal organization and dynamics in synthetic cells
A. Groaz, H. Moghimianavval, F. Tavella, T. Giessen, A. Vecchiarelli, Q. Yang, A. Liu
WIREs Nanomedicine and Nanobiotechnology (2020)
DOI: 10.1002/wnan.1685 | Link
A. Groaz, H. Moghimianavval, F. Tavella, T. Giessen, A. Vecchiarelli, Q. Yang, A. Liu
WIREs Nanomedicine and Nanobiotechnology (2020)
DOI: 10.1002/wnan.1685 | Link
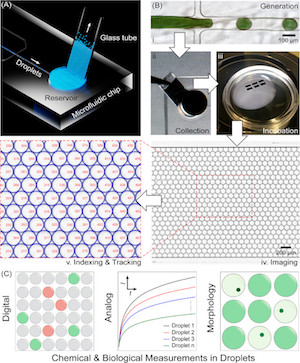
Building dynamic cellular machineries in droplet-based artificial cells with single-droplet tracking and analysis
M. Sun, Z. Li, S. Wang, G. Maryu, and Q. Yang
Analytical Chemistry (2019)
DOI: 10.1021/acs.analchem.9b01481 | Link
M. Sun, Z. Li, S. Wang, G. Maryu, and Q. Yang
Analytical Chemistry (2019)
DOI: 10.1021/acs.analchem.9b01481 | Link

μdroPi: a hand-held microfluidic droplet imager and analyzer built on Raspberry Pi
M. Sun, Z. Li, and Q. Yang
Journal of Chemical Education (2019)
DOI: 10.1021/acs.jchemed.8b00975 | Link
M. Sun, Z. Li, and Q. Yang
Journal of Chemical Education (2019)
DOI: 10.1021/acs.jchemed.8b00975 | Link
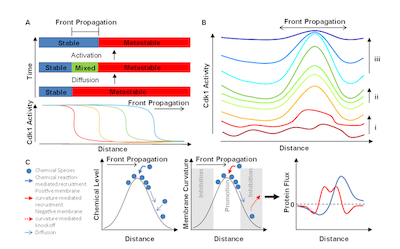
The rise of ultrafast waves
O. Puls and Q. Yang
Developmental Cell (2018)
DOI: 10.1016/j.devcel.2018.11.026 | Link
O. Puls and Q. Yang
Developmental Cell (2018)
DOI: 10.1016/j.devcel.2018.11.026 | Link
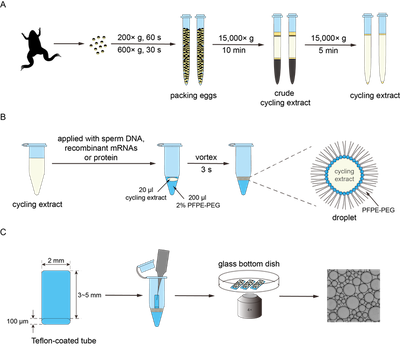
Reconstitution of cell-cycle oscillations in microemulsions of cell-free Xenopus egg extracts
Y. Guan, S. Wang, M. Jin, H. Xu, and Q. Yang
Journal of Visualized Experiments (2018)
DOI: 10.3791/58240 | Link
Y. Guan, S. Wang, M. Jin, H. Xu, and Q. Yang
Journal of Visualized Experiments (2018)
DOI: 10.3791/58240 | Link
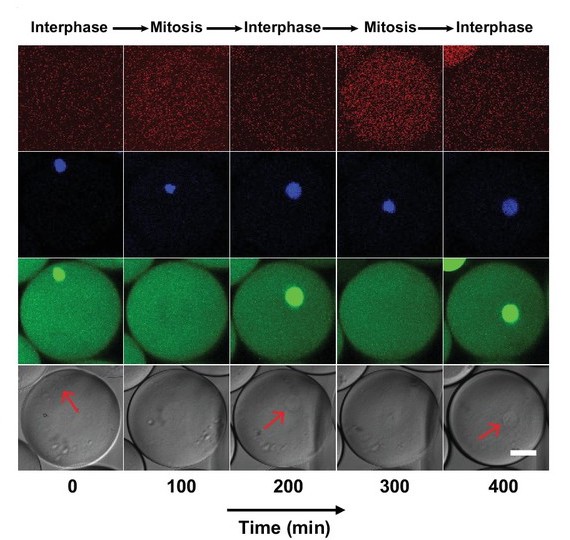
A robust and tunable mitotic oscillator in artificial cells
Y. Guan*, Z. Li*, S. Wang, P. Barnes, X. Liu, H. Xu, M. Jin, A.P. Liu, and Q. Yang
eLife (2018)
DOI: 10.7554/eLife.33549 | Link
Y. Guan*, Z. Li*, S. Wang, P. Barnes, X. Liu, H. Xu, M. Jin, A.P. Liu, and Q. Yang
eLife (2018)
DOI: 10.7554/eLife.33549 | Link
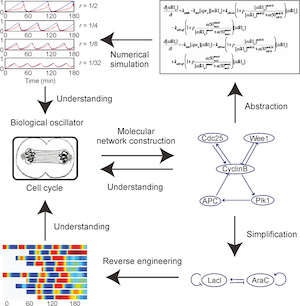
Systems and synthetic biology approaches in understanding biological oscillators
Z. Li and Q. Yang
Quantitative Biology (2018)
DOI: 10.1007/s40484-017-0120-7 | Link
Z. Li and Q. Yang
Quantitative Biology (2018)
DOI: 10.1007/s40484-017-0120-7 | Link
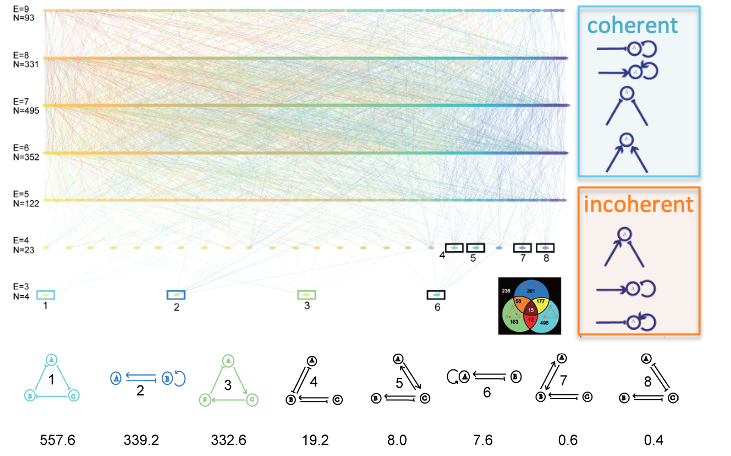
Incoherent inputs enhance the robustness of biological oscillators
Z. Li, S. Liu, and Q. Yang
Cell Systems (2017)
DOI: 10.1016/j.cels.2017.06.013 | Link
Z. Li, S. Liu, and Q. Yang
Cell Systems (2017)
DOI: 10.1016/j.cels.2017.06.013 | Link
2014 and Earlier¶
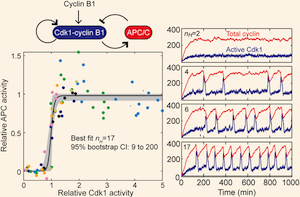
The Cdk1-APC/C cell cycle oscillator circuit functions as a timedelayed, ultrasensitive switch
Q. Yang and J. E. Ferrell Jr
Nature Cell Biology (2013)
DOI: 10.1038/ncb2737 | Link
Q. Yang and J. E. Ferrell Jr
Nature Cell Biology (2013)
DOI: 10.1038/ncb2737 | Link
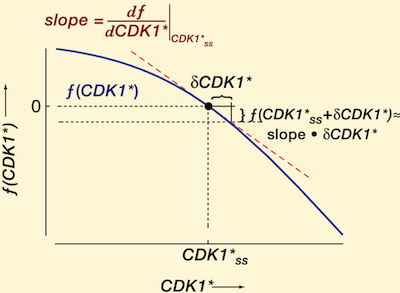
Modeling the cell cycle: why do certain circuits oscillate?
J. E. Ferrell Jr., T. Tsai, and Q. Yang
Cell (2011)
DOI: 10.1016/j.cell.2011.03.006 | Link
J. E. Ferrell Jr., T. Tsai, and Q. Yang
Cell (2011)
DOI: 10.1016/j.cell.2011.03.006 | Link
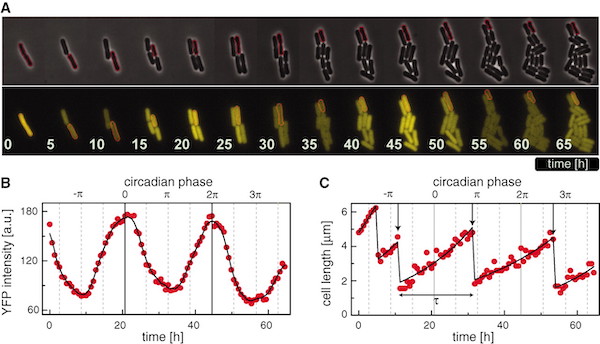
Circadian gating of the cell cycle revealed in single cyanobacterial cells
Q. Yang*, B. F. Pando*, G. Dong, S. S. Golden, and A. van Oudenaarden
Science (2010)
DOI: 10.1126/science.1181759 | Link
Q. Yang*, B. F. Pando*, G. Dong, S. S. Golden, and A. van Oudenaarden
Science (2010)
DOI: 10.1126/science.1181759 | Link
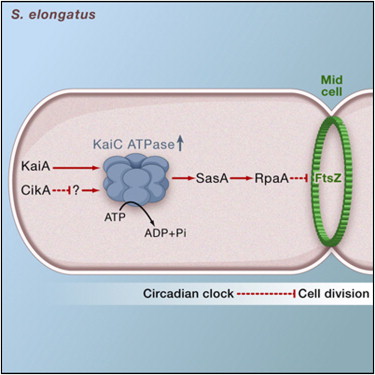
Elevated ATPase activity of KaiC applies a circadian checkpoint on cell division in Synechococcus elongatus
G. Dong, Q. Yang, Q. Wang, Y. Kim, T. L. Wood, K. W. Osteryoung, A. van Oudenaarden, and S. S. Golden
Cell (2010)
DOI: 10.1016/j.cell.2009.12.042 | Link
G. Dong, Q. Yang, Q. Wang, Y. Kim, T. L. Wood, K. W. Osteryoung, A. van Oudenaarden, and S. S. Golden
Cell (2010)
DOI: 10.1016/j.cell.2009.12.042 | Link
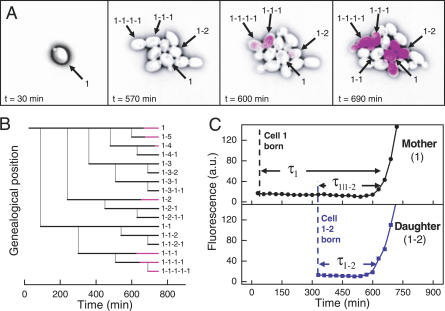
Heritable stochastic switching revealed by single-cell genealogy
B. B. Kaufmann*, Q. Yang*, J. T. Mettetal, and A. van Oudenaarden
PLoS Biology (2007)
DOI: 10.1371/journal.pbio.0050239 | Link
B. B. Kaufmann*, Q. Yang*, J. T. Mettetal, and A. van Oudenaarden
PLoS Biology (2007)
DOI: 10.1371/journal.pbio.0050239 | Link
Software Developed in the Yang Lab¶
First plot: — B nullcline — C nullcline — Trajectory
Second plot: — Total Cyclin B1 (B) — Active Cdk1:Cyclin B1 (C)
Second plot: — Total Cyclin B1 (B) — Active Cdk1:Cyclin B1 (C)
Credit: Interactive Cell Cycle Model - Yang Lab by Franco Tavella